Note
Go to the end to download the full example code.
Regression Conformal Prediction — sklearn¶
Demonstrate absolute_error_score()
using a DecisionTreeRegressor on the Diabetes dataset.
The conformal interval for a new point \(x\) is
\[\hat{f}(x) \pm \hat{q}_{1-\alpha}\]
where \(\hat{q}_{1-\alpha}\) is the empirical \((1-\alpha)\)-quantile of the absolute residuals on the calibration set.
from __future__ import annotations
import matplotlib.pyplot as plt
import numpy as np
from sklearn.datasets import load_diabetes
from sklearn.ensemble import RandomForestRegressor
from sklearn.model_selection import KFold, train_test_split
from probly.calibrator import calibrate
from probly.metrics._common import average_interval_size, empirical_coverage_regression
from probly.method.conformal import conformal_absolute_error
from probly.representer import representer
Data preparation¶
Build and train the model¶
model = RandomForestRegressor(random_state=42)
model.fit(X_train, y_train)
Absolute error score¶
calibrated_model = calibrate(conformal_absolute_error(model), 0.05, y_calib, X_calib)
output = representer(calibrated_model).predict(X_test)
coverage = empirical_coverage_regression(output, y_test)
avg_size = average_interval_size(output)
print(f"Absolute Error — coverage: {coverage:.3f}, avg interval size: {avg_size:.1f}")
Absolute Error — coverage: 0.955, avg interval size: 209.7
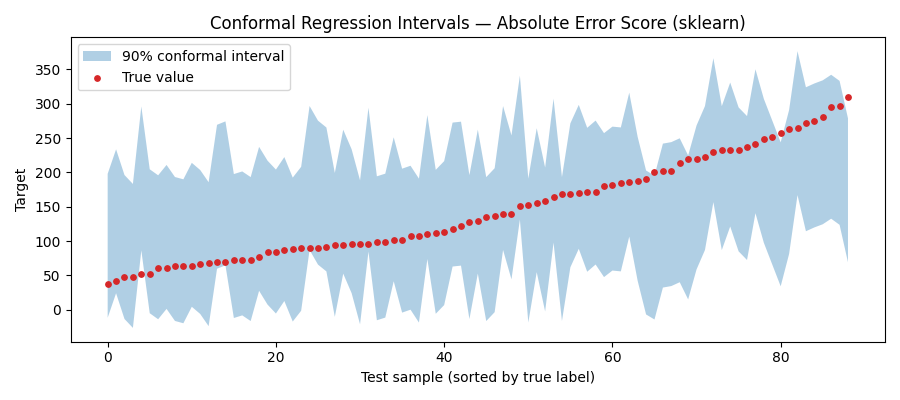
Visualise prediction intervals¶
Sort by true label for a cleaner plot.
intervals = output.array # (n_test, 2): [lower, upper]
order = np.argsort(y_test)
plt.figure(figsize=(9, 4))
plt.fill_between(
range(len(y_test)),
intervals[order, 0],
intervals[order, 1],
alpha=0.35,
label="90% conformal interval",
)
plt.scatter(range(len(y_test)), y_test[order], s=15, color="tab:red", label="True value", zorder=3)
plt.xlabel("Test sample (sorted by true label)")
plt.ylabel("Target")
plt.title("Conformal Regression Intervals — Absolute Error Score (sklearn)")
plt.legend()
plt.tight_layout()
plt.show()
Summary (Averaged over multiple runs)¶
res = {"Absolute Error": []}
for fold, (train_idx, test_idx) in enumerate(KFold(n_splits=5, shuffle=True, random_state=42).split(X)):
X_train, y_train = X[train_idx], y[train_idx]
X_test, y_test = X[test_idx], y[test_idx]
X_train, X_calib, y_train, y_calib = train_test_split(X_train, y_train, test_size=0.25, random_state=fold)
fold_model = RandomForestRegressor(max_depth=3, random_state=fold)
fold_model.fit(X_train, y_train)
calibrated_model = calibrate(conformal_absolute_error(fold_model), 0.05, y_calib, X_calib)
output = representer(calibrated_model).predict(X_test)
cov = empirical_coverage_regression(output, y_test)
size = average_interval_size(output)
res["Absolute Error"].append((cov, size))
for name, vals in res.items():
covs, sizes = zip(*vals)
print(f"{name} — coverage: {np.mean(covs):.3f} ± {np.std(covs):.3f}, avg interval size: {np.mean(sizes):.1f} ± {np.std(sizes):.1f}")
Absolute Error — coverage: 0.959 ± 0.027, avg interval size: 238.1 ± 15.4
Total running time of the script: (0 minutes 0.965 seconds)